So, there are two main objectives of this script: display data based on a certain threshold value (in this case, reflectance < 1000) and make a standard deviation from the results. The Data used in this example was Sentinel-2 MSI Level 2A, in the period between November 1 - 15, 2019. Cloud masking was utilized at the beginning of the script to choose granules which were clear from clouds.
// Create a geometry representing the analysis region.
var daerah = ee.Geometry.Rectangle([72.611536, 28.354144, 78.807625, 32.630466]);
/**
* Function to mask clouds using the Sentinel-2 QA band
* @param {ee.Image} image Sentinel-2 image
* @return {ee.Image} cloud masked Sentinel-2 image
*/
function maskS2clouds(image) {
var qa = image.select('QA60');
// Bits 10 and 11 are clouds and cirrus, respectively.
var cloudBitMask = 1 << 10;
var cirrusBitMask = 1 << 11;
// Both flags should be set to zero, indicating clear conditions.
var mask = qa.bitwiseAnd(cloudBitMask).eq(0)
.and(qa.bitwiseAnd(cirrusBitMask).eq(0)
// Choose pixels with reflectance < 1000
.and(image.lt(1000))
);
return image.updateMask(mask).divide(10000);
}
// Map the function over the analysis region and period
// Load Sentinel-2 TOA reflectance data.
var datasentinel = ee.ImageCollection('COPERNICUS/S2')
.filterDate('2019-11-01', '2019-11-15')
.filterBounds(daerah)
// Pre-filter to get less cloudy granules.
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 20))
.map(maskS2clouds)
;
// Get the standard Deviation.
var hasil = datasentinel.reduce(ee.Reducer.stdDev());
// Define the plot parameter, use the standard deviation of Band 4 only
var stdevVis = {
min: 0,
max: 0.005,
bands: ['B4_stdDev'],
palette: ['0000FF', '00FFFF', '00FF00', 'FFFF00', 'FF0000']
};
// Add the plot layer on the map
Map.addLayer(hasil, stdevVis, 'Std Deviation');
Map.setCenter(75.7095805, 30.492305, 14);
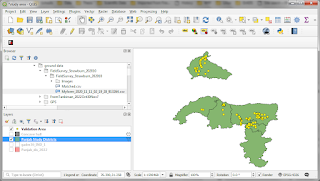
And the result will be like this:
You may seen some pixels with very high standard deviation values (shown in red, cyan or anything else other than blue). To 'clean-up' these pixels, we can add another filter to the standard deviation results. For example to show the values no higher than 0.000001:
// Get the standard Deviation.
var hasil = datasentinel.reduce(ee.Reducer.stdDev());
var hi_stddev = hasil.lt(0.000001);
var hasil = hasil.updateMask(hi_stddev);
The result will be like this:
Finally, we can overlay the base Google Map using the result by choosing the Satellite map (button on the right corner of the map).